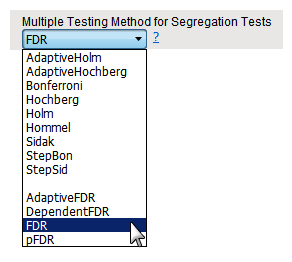
This drop-down menu enables you to specify a method for adjusting for multiple hypothesis tests across all segregation ratios.
The AdaptiveHolm, AdaptiveHochberg, Bonferroni, Hochberg, Holm, Hommel, Sidak, StepBon, and StepSid methods all control for the familywise error rate. The methods based on FDR all control for false discovery rate.
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
|||||
|
Hochberg, Y. and Benjamini, Y. (1990). More Powerful Procedures for Multiple Significance Testing. Statistics in Medicine 9: 811–818.
Hochberg, Y. (1988). A Sharper Bonferroni Procedure for Multiple Significance Testing. Biometrika 75: 800–803.
Hommel, G. (1988). A Comparison of Two Modified Bonferroni Procedures. Biometrika 75: 383–386.
Benjamini, Y. and Hochberg, Y. (2000). On the Adaptive Control of the False Discovery Rate in Multiple Testing with Independent Statistics. Journal of Educational and Behavioral Statistics 25: 60–83.
Benjamini, Y. and Yekateuli, D. (2001). The Control of the False Discovery Rate in Multiple Testing under Dependency. Annals of Statistics 29: 1165–1188.
Benjamini, Y. and Hochberg, Y. (1995). Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society, B 57: 289–300.
Storey JD. (2002) A direct approach to false discovery rates. Journal of the Royal Statistical Society, Series B, 64: 479-498.
Storey JD, Taylor JE, and Siegmund D. (2004). Strong control, conservative point estimation, and simultaneous conservative consistency of false discovery rates: A unified approach. Journal of the Royal Statistical Society, Series B, 66: 187-205.
Refer to the SAS PROC MULTTEST documentation for more details about each of these methods.