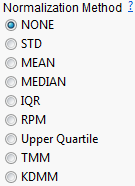
Select a normalization method.
|
|||||||
|
|||||||
|
|||||||
|
|||||||
|
|||||||
|
|||||||
|
|||||||
|
|||||||
|
|||||||
For more information, refer to the Data Standardize process description.