The Data Standardize normalization process standardizes values of numeric variables in a SAS data set. A large number of potential methods are available for performing the standardization.
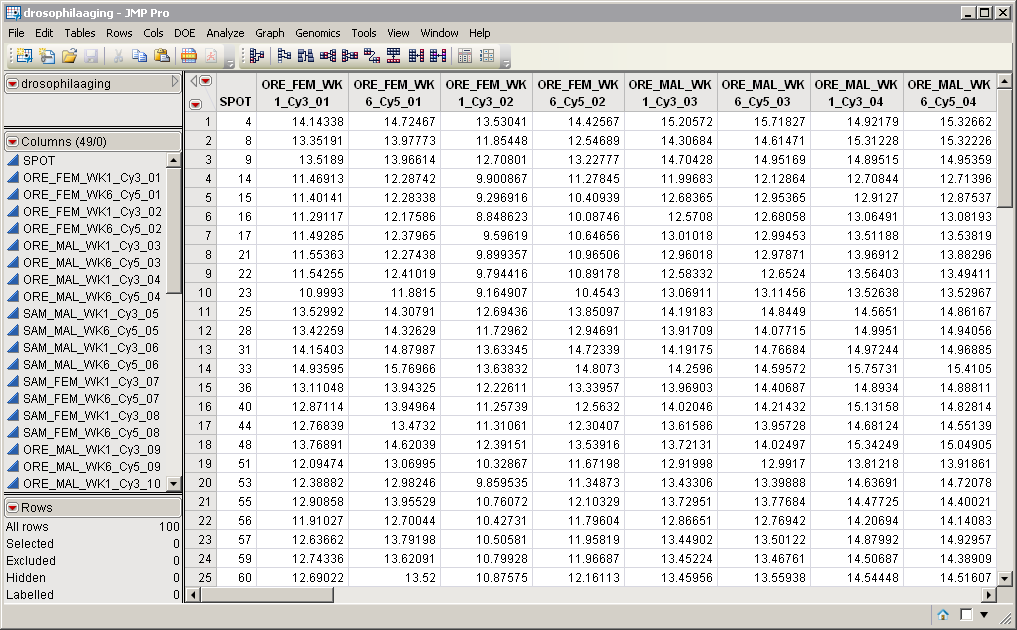
One data set is required for this process: the Input Data Set that contains all of the numeric data to be analyzed. The drosophilaaging.sas7bdat data set from the Drosophila aging experiment of Jin, et al. (2001) that is described in Drosophila Aging Experimental Data and used in the example that follows, is shown below (and included in the Sample Data folder that comes with JMP Genomics). It has 48 data columns and 100 rows. Note that this is a tall data set; each probe corresponds to one row whereas each column corresponds to a separate experimental condition.
|
•
|
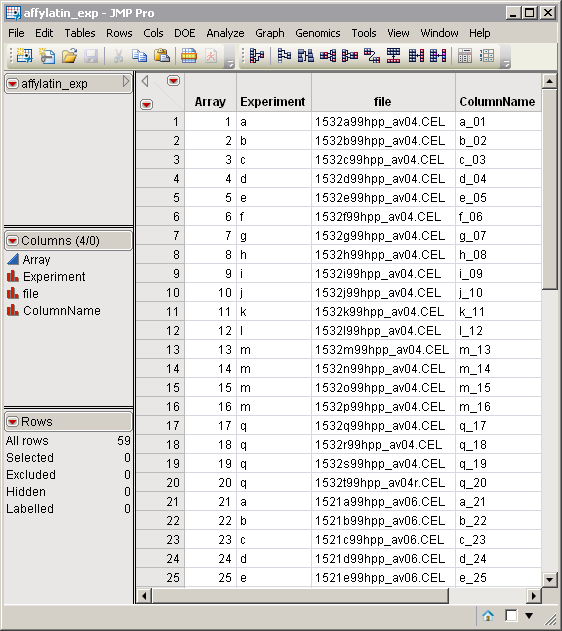
The Experimental Design Data Set (EDDS). The drosophilaaging_exp.sas7bdat EDDS is shown below. This data set tells how the experiment was performed, providing information about the columns in the input data set. Note that one column in the EDDS must be named ColumnName and the values contained in this column must exactly match the column names in the input data set.
|
|
•
|
The Standardization Statistics Input SAS Data Set (not shown). This data set is used when selecting DATASET as the Standardization Method. It contains location and scale estimates by which to standardize the input data set.
|
|
•
|
The Drosophila aging experiment of Jin, et al. (2001) consists of 24 two-color cDNA microarrays, six for each experimental combination of two lines (Oregon and Samarkand), two sexes (Female and Male), and two ages (1 week and 6 weeks). The Cy3 and Cy5 dyes were flipped for two of the six replicates for each genotype and sex combination. The design is a split-plot design, with Age and Dye as subplot factors, and Line and Sex as whole-plot factors. A total of 4256 clones were spotted on the arrays, but for this example, we use a subset containing 100 randomly selected genes.
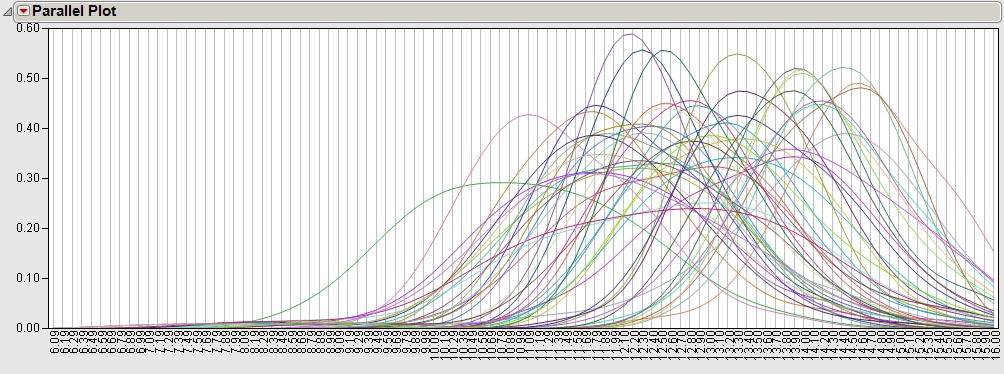
The raw data from this experiment were evaluated using distribution analysis. The Parallel Plot (shown below) provides the raw univariate distributions of all 48 channels from the 24 arrays.
|
|
Select MEDIAN as the standardization method to follow this example.
|
For detailed information about the files and data sets used or created by JMP Life Sciences software, see Files and Data Sets.
Refer to the Data Standardize output documentation for detailed descriptions of the output of this process.