Process Description
Correlation and Principal Variance Component Analysis
The Correlation and Principal Component process computes correlations between numeric variables, principal components of the correlation matrix and an accompanying outlier analysis. It also optionally computes a variance components decomposition that helps you determine the major sources of overall variability in your experiment.
What do I need?
Two data sets are required for this process.
The first data set, the Input Data Set, contains all of the numeric data to be analyzed. This data set must be in the tall format where each sample corresponds to one row and each column corresponds to a separate experimental condition or array.
The drosophilaaging_norm.sas7bdat data set, shown below, is a normalized data set derived from the Drosophila Aging experiment described in Sample Case Studies. It has 49 columns and 100 rows corresponding to 49 arrays and 100 individual probes, respectively.
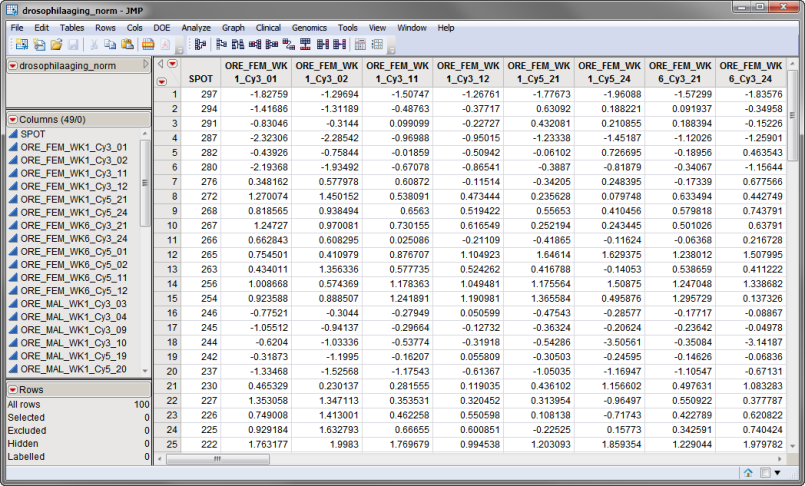
The second data set is the Experimental Design Data Set (EDDS). This required data set tells how the experiment was performed, providing information about the columns in the input data set. Note that one column in the EDDS must be named ColumnName and the values contained in this column must exactly match the column names in the input data set.
The drosophilaaging_exp.sas7bdat EDDS, is shown below. Note that the ColumnName column lists the column names in the input data set. The Array column corresponds to an index variable. Note the variables describing experimental conditions.

For detailed information about the files and data sets used or created by JMP Genomics software, see Files and Data Sets.
Output/Results
The output generated by this process is summarized in a Tabbed report. Refer to the Correlation and Principal Variance Component Analysis output documentation for detailed descriptions and guides to interpreting your results.