Principal components
analysis (PCA) is a useful tool for exploring and revealing population structure based on SNP
genotypes
for a sample. It can also be used to adjust for
population stratification
and
allele
frequency variation due to ancestral differences, in
association
tests between
SNP
genotypes and a
binary
or
continuous trait
via the EIGENSTRAT method (Price
et al
., 2006).
The
PCA for Population Stratification
process performs PCA on the rows (individuals) of the input data set to infer axes of genetic variation and adjust the association test accordingly. It produces several plots (like those created by the
Principal Components Analysis
process), and also outputs
p-value
plots like those created by the other Association Testing processes, that include the EIGENSTRAT adjustment for population stratification.
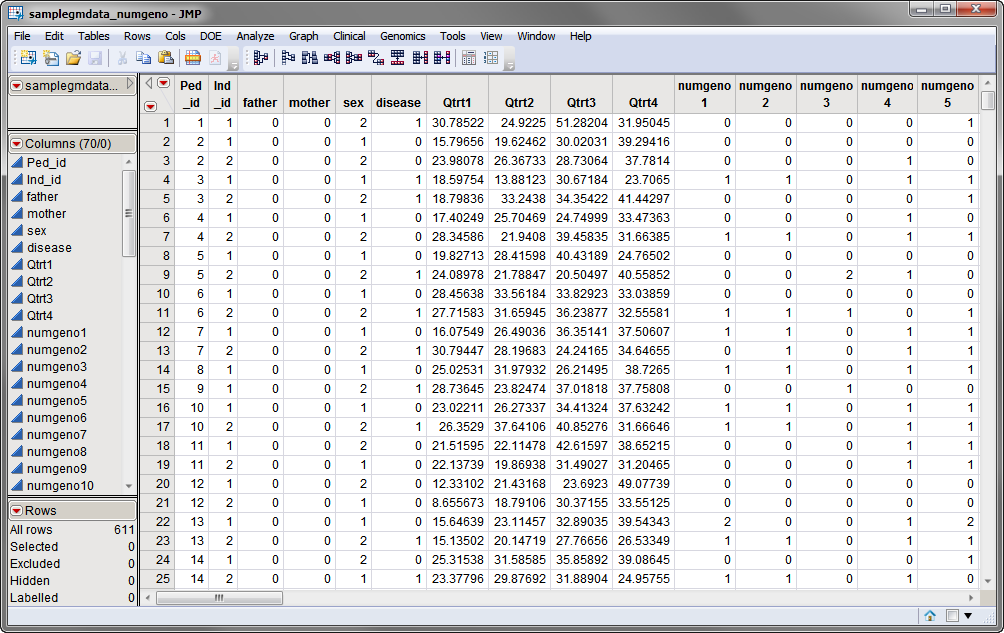
Two data sets are needed to run this process. The first, the
Input Data Set
, contains all of the marker data. The sample data set used in the following example, the
samplegmdata_numgeno
data set, is partially shown below.
The original
samplegmdata
data set described in
Data Sets Used in JMP Genomics Processes
was computer generated and consists of 1000 rows of individuals with 130 columns corresponding to data on these individuals. Marker data is presented in the two-column
allelic
format. This data set was recoded using the
Recode Genotypes
process to generate
samplegmdata_numgeno
data set. The recoded data set consists of 611 rows with 70 columns. Marker data is presented as
numeric variables
in the one-column genotypic format.
Note
:
The
marker variables
in the input data set must contain numerically coded genotypes; a data set containing these can be obtained by checking the
Create data set with numerically coded genotypes
option when you run
Marker Properties
or by running the
Recode Genotypes
process.
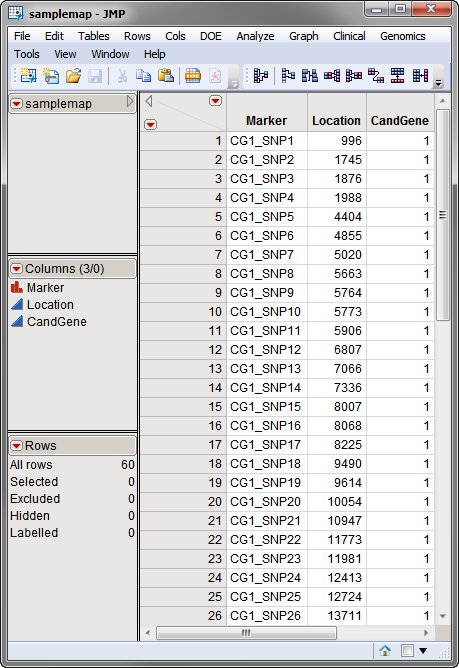
The second data set is the
Annotation Data Set
. This optional data set contains information, such as gene identity or chromosomal location, for each of the markers. The
annotation data set
used in this example, the
samplemap
data set, was computer generated and identifies markers, location and gene identities. A portion of this data set is illustrated below. This data set is a tall data set; each row corresponds to a different marker.
Both the
samplegmdata
and
samplemap
data sets are included in the
Sample Data
folder that comes with JMP Genomics.
For detailed information about the files and data sets used or created by JMP Life Sciences software, see
Files and Data Sets
.
The output generated by this process is summarized in a Tabbed report. Refer to the
PCA for Population Stratification
output documentation for detailed descriptions and guides to interpreting your results.