Unlike normal
microarrays
, which typically include a limited number of spaced, non-overlapping,
probes
for each known gene,
tiling
arrays contain many overlapping probes designed to cover large contigs of the
genome
, or even the whole genome itself. Because of their potential for very high resolution, and coverage, tiling arrays can produce a much more unbiased examination of overall
expression
of both known and unidentified genes. However, because of the immense number of probes, the
.cel
files containing the data tend to be very large.
The
Basic Tiling Workflow
enables you to run a basic
workflow
for tiling data. The workflow includes options for quality control and
ANOVA
.
|
•
|
An
Input Data Set
. Before you can run the
Basic Tiling
Workflow, you must import the data into SAS data sets from the raw data files. The
Affymetrix Tiling CEL Input Engine
can be used to import the data from Affymetrix
.cel
files into an
input SAS data set
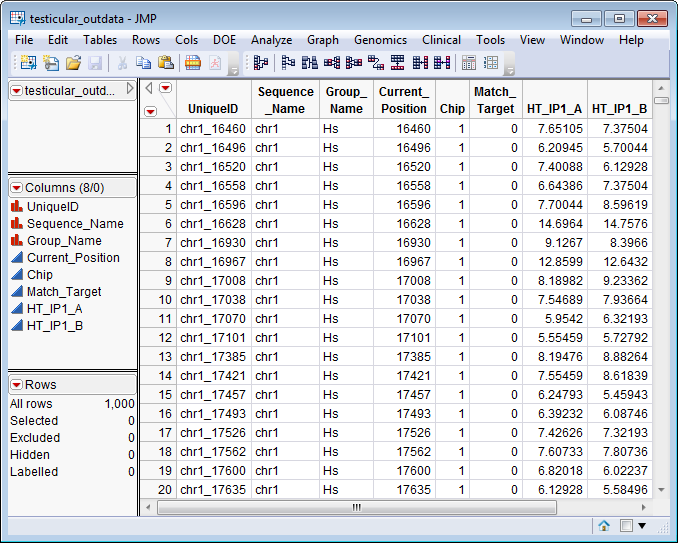
. The
testicular_outdata.sas7bdat
data set serves as an example, and is shown below.
|
|
•
|
An
Experimental Design Data Set (EDDS)
for the experiment. The EDDS lists specific information about the design of the experiment. The EDDS is a SAS data set that must be created before the data can be imported. An EDDS is created by the
Affymetrix Tiling CEL Input Engine
during data import
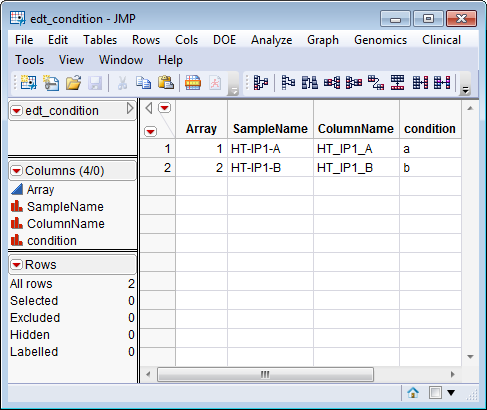
. The
edt_condition.sas7bdat
EDDS serves as an example, and is shown below.
|
A third data set, the
Annotation Data Set
, is
optional
. This data set contains information such as gene identity or chromosomal location, for each of the markers.
For detailed information about the files and data sets used or created by JMP Life Sciences software, see
Files and Data Sets
.
When you click
, the
Basic Tiling Workflow
process begins by opening the
Workflow Builder
. The
Workflow Builder
builds
settings files
for each process, containing the information from the data sets and parameters specified in the
Basic Tiling Workflow
dialog
. Once the setting files are generated and saved, the individual processes in the workflow are sequentially opened, populated, and run. The results of the processes are saved in the specified output folder. Finally, a JMP
journal
is generated, providing links to the workflow dialog and the results of each process.
|
|
Click
.
|
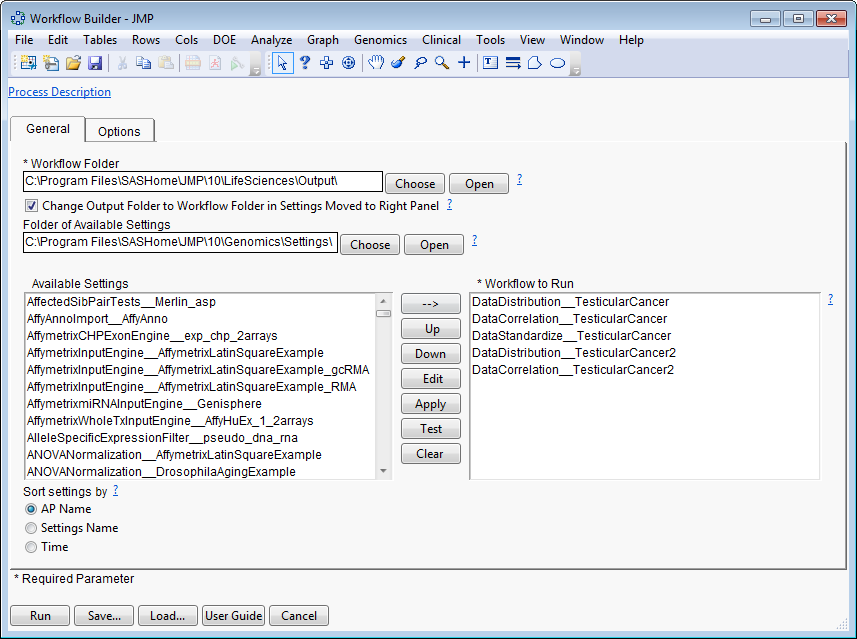
The
Workflow Builder
dialog shows the settings for each of the processes in the workflow. You can select and edit individual settings to adjust your analysis.
|
|
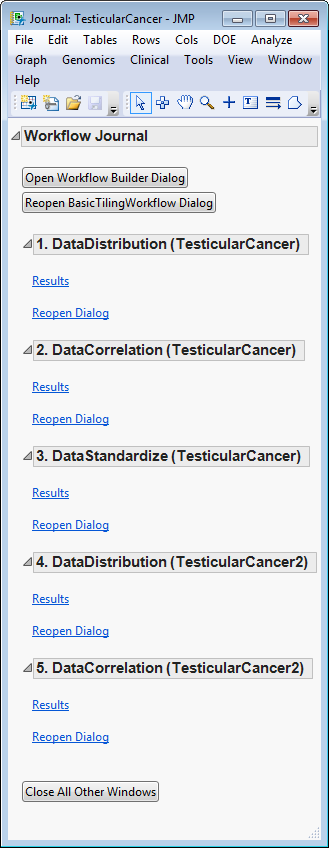
Clicking each of the
buttons on the journal brings up the output of each of the processes. This enables you to examine each set of output so that adjustments can be made to the individual settings, as needed. For your convenience, links to the default
Basic Tiling Workflow
processes are given below.
|