The
Chromosome Results
tab is shown below:
The
Chromosome Results
tab contains the following elements:
|
•
|
One
P
-Value Plot.
|
There is a separate results tab for each
chromosome
containing one
p-value
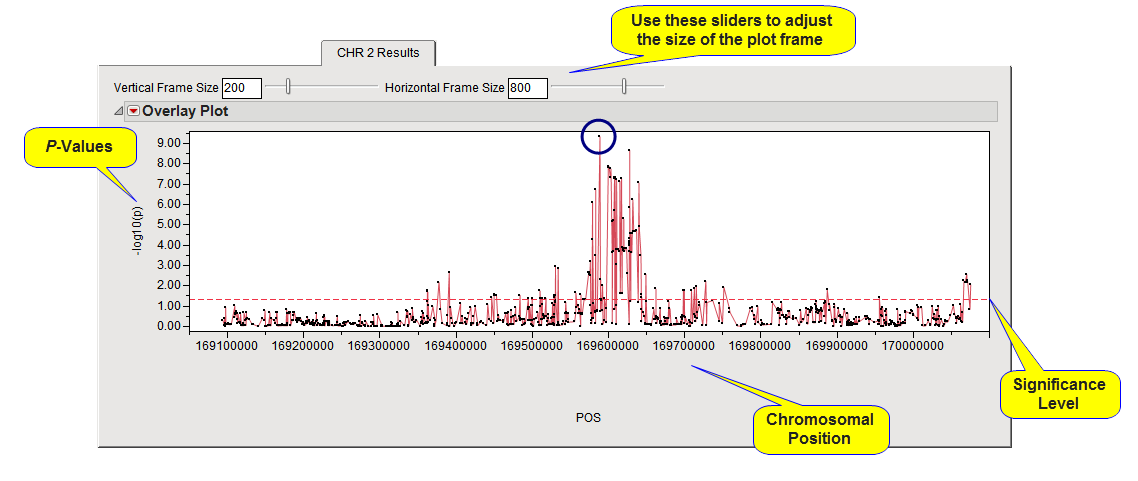
plot when multiple chromosomes are included in the analysis. The tab for chromosome 2 is shown above.
The
Y
-axis
variable
is the meta-analysis
p
-value, converted to the
-log
or
-log
10
scale if selected in the process
dialog
. The
X
-axis plots location of the marker according to the
Location Variable
within the chromosome.
A horizontal reference line is drawn as a red, dashed line at the significance level that was specified. For
-log
or
-log
10
-converted
p
-values, markers above this line are significant; for
p
-values on the original scale, markers below the line are significant. The marker showing the highest significance (SNP rs560887) is circled in the plot above.
On this plot or any of the other
p
-value plots, simply mouse-over any of the points on the plot to see the marker's label, for example, reference
SNP
ID. When an annotation accession variable is specified when running the process, you can select a point and click on any of the Annotation action buttons (
GenBank Nucleotide
,
UniGene Database
,
AceView Database
, or
dbSNP
) to link directly to the corresponding website to view extensive annotation information about the
locus
.
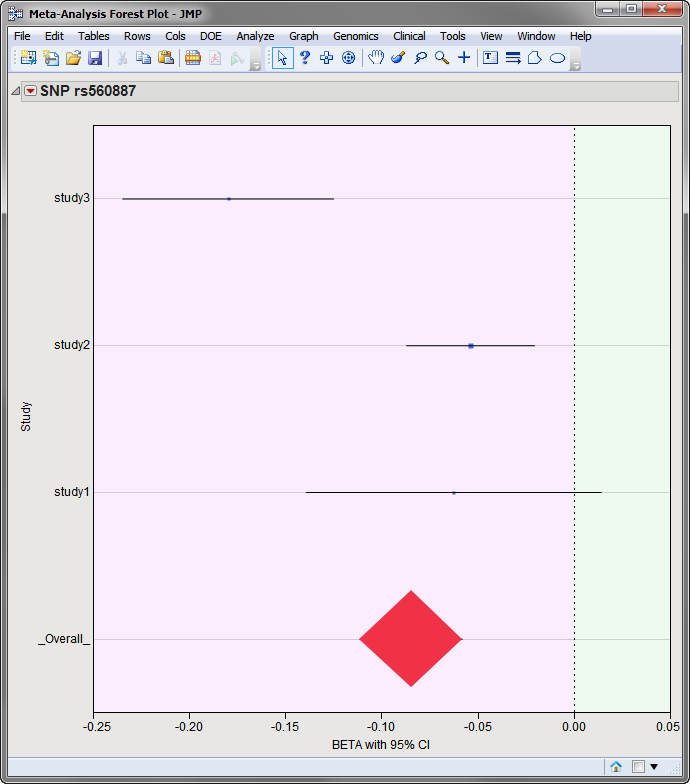
Also, on this plot or any of the other
p
-value or heterogeneity plots, you can select markers for which to create
forest plots
by clicking on the
Forest Plot
action button. A separate forest plot is created for each selected marker showing the effect estimate (
regression
coefficient or
log odds
ratio, for example) for each individual study along with its confidence interval, and the weight for the study (in this case, the inverse of the effect estimate's
variance
) is represented by the size of the square. A red diamond is centered at the overall estimate with width extending the length of the confidence interval, as shown for marker SNP rs560887 (circled above) below:
Note
: The analysis used in this example consisted of three separate studies.