The
Linkage Map
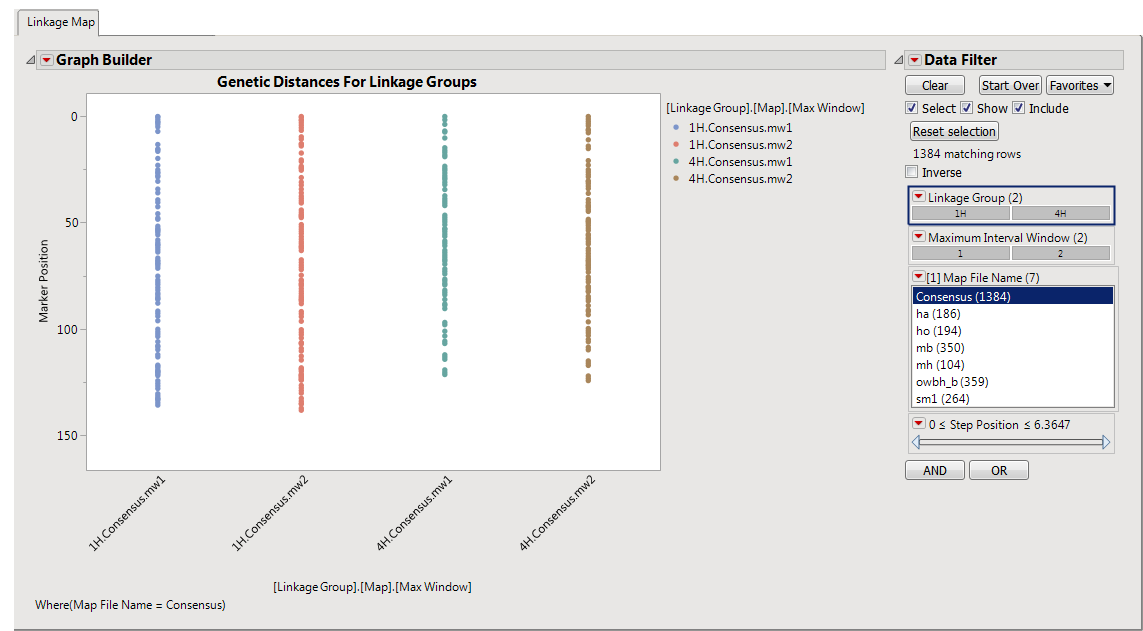
tab is shown below:
|
•
|
One
Genetic Distances for Linkage Groups
plot.
|
This plot displays the position of markers (
y
-axis) versus the chromosome index (
x
-axis) for each chromosome. Each colored filled-circle represents a marker on the chromosome.
A chromosome index for the consensus map is defined as a concatenation of the input linkage group name, a dot, the word
Consensus
(if the map is a consensus output), a dot, and the maximum interval window used to build the consensus map (if the map is a consensus output). For example, if the input linkage group name is
1H
and the map is a consensus output build with maximum interval window of 2, then the chromosome index is
1H.Consensus.mw2
.
Chromosome indexes for the original input linkage groups are likewise defined, however, with the word
Consensus
replaced by the name of the input map file, and without maximum interval window. For example, if an input linkage group has name
1H
, and it is coming from an input file named
mb.sas7bdat
, then its chromosome index is
1H.mb
.
Select points in one or more chromosomes and click
to launch the
Linkage Map Viewer
process to visualize the selected chromosomes. Likewise, select points in two or more chromosomes and click
to launch the
Compare Linkage Maps
process to compare all pairwise selected chromosomes.
The JMP
Data Filter
has also been embedded into this tabbed view to use for interactively selecting points in the plots.