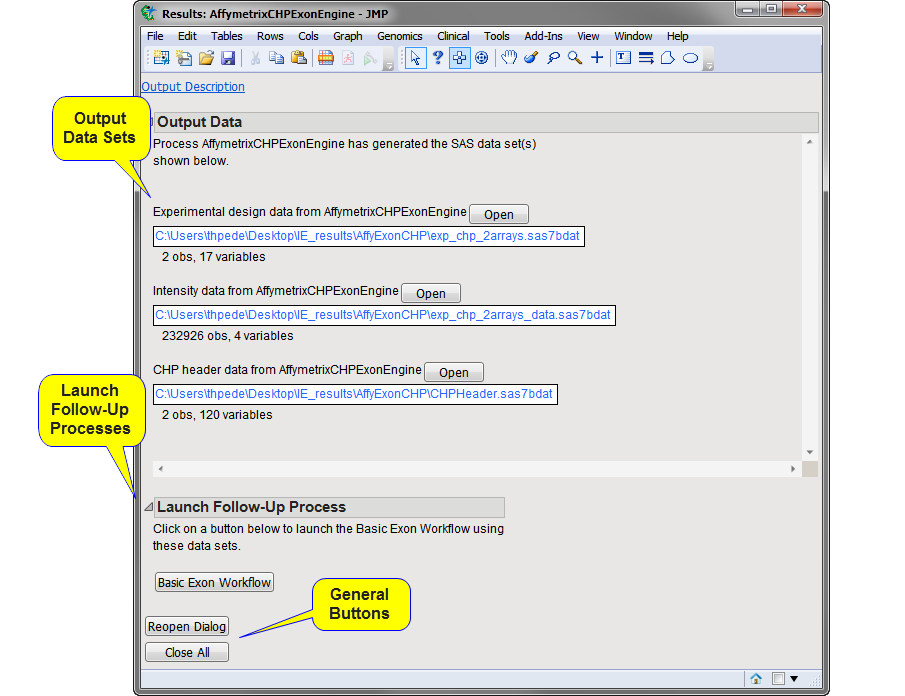
Running this process for the Affymetrix Human Exon 1.0 ST colon cancer data set generates the
Results
window shown below. Refer to the
Affymetrix Exon CHP Input Engine
process description for more information.
The
Results
window contains the following elements:
|
•
|
Experimental Design Data Set (EDDS)
: This data set lists specific information about experimental details for each of the arrays. Refer to
Experimental Design Data Set (EDDS)
for more information about this data set.
|
|
•
|
Output Data Set
: This data set contains the data from the different
.chp
files. This data set is in the tall format: the intensity data from each array are listed in a separate column while individual spots are listed in rows.
|
|
•
|
CHP Header Data Set
: This data set contains all the parameters in the header portion of each Command Console file. These parameters include the algorithm applied, the type of chip used, the program applied, and other quality control information among the control l
probes
. The parameters are listed in columns, while the individual
.cel
files are listed in rows.
|
|
•
|
Quality Flag Data Set
: One or more of these data sets are generated when the .
chp
files, such as MAS5 CHP files that contain a
Detection
column, are imported. The
QualityFlag
data set contains columns with quality information about the intensity of each
probeset
for each sample. In this situation, a column called
Detection_Percentage
, containing numeric codes (
0
=present,
1
=marginal, and
2
=absent), is generated in both the
QualityFlag
data set and the main output data table. You can use this parameter, which represents the percentage of present calls for that row (
probeset
) across all arrays in the data set, to filter out genes with low or marginal signals from the analysis.
Note
: This data set is accessed from the specified
Output Folder
.
|
Note
: You can view the data in a file or data set by clicking either
or
(when the data set is very large).
Tip
: To filter the
Quality Flag
data set directly using the
Detection_Percentage
column, use the
Filter Intensities
process. Replace all values in the
Detection_Percentage
column falling below an acceptable percentage with the
missing value
symbol (
.
) and then delete all rows with a
missing value
symbol. Merge the filtered
QualityFlag
data set back to the main data table, using the
Merge
process.
|
•
|
Basic Exon Workflow
: Click
to launch the
Basic Exon/Alternative Splicing Workflow
process with the output data set and the EDDS specified as input.
|
|
•
|
Click
to reopen the completed process dialog used to generate this output.
|
|
•
|
Click
to close all graphics windows and underlying data sets associated with the output.
|