The
Missing Genotype by Trait Summary
process tests whether there are significant differences in missing
genotype
proportions between cases and controls (or any two groups as defined by a
binary trait
variable) for each individual marker. The
p-values
from these tests are displayed in a plot along the marker map as are the number of missing genotypes in each
trait
group.
Distributions
for the number of missing genotypes across all markers are also displayed for each trait group.
The
Missing Genotype by Trait Summary
process tests whether missing genotypes at a particular marker are related to a trait. It should therefore be run before running any
association
testing.
Two data sets are needed to run the
Missing Genotype by Trait Summary
process. The first, the
Input Data Set
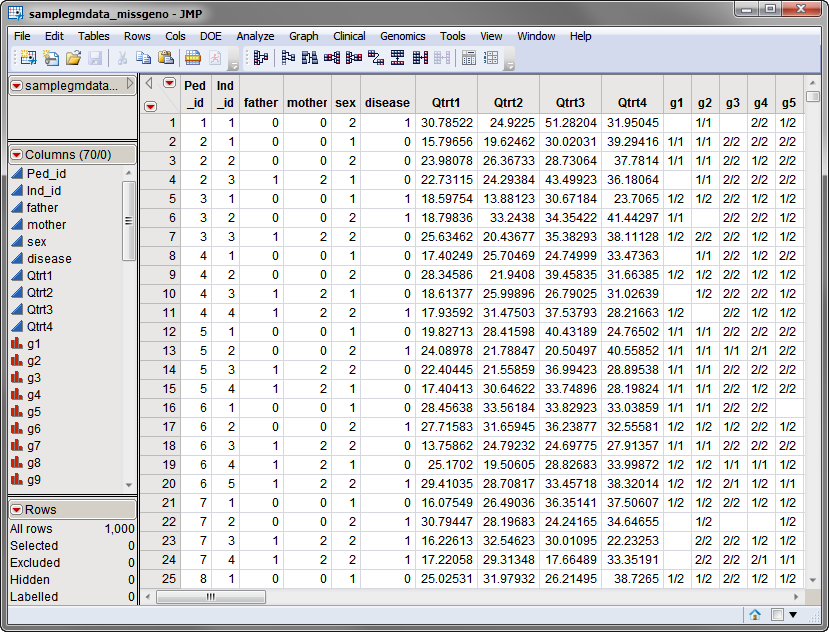
, contains all of the marker data. The
samplegmdata_missgeno.sas7bdat
data set used in the following example was generated from the
samplegmdata.sas7bdat
data set described in
Data Sets Used in JMP Genomics Processes
. This data set was computer generated and consists of 1000 rows of individuals with 130 columns corresponding to data on these individuals. Data for 60 markers is presented in the two-column
allelic
format in columns
ma1
-
ma120
. Modifications, found in the
samplegmdata_missgeno.sas7bdat
data set, include the conversion of the marker data from the two-column allelic format to the one column
genotypic
format (60 columns,
g1
-
g60
), in which the
alleles
are delimited by a
/
and selective elimination of data at different markers in different individuals. This data set is partially shown below. Note that this is a wide data set; markers are listed in columns, whereas individuals are listed in rows.
The second optional data set is the
Annotation Data Set
. This data set contains information, such as gene identity or chromosomal location, for each of the markers. The
annotation data set
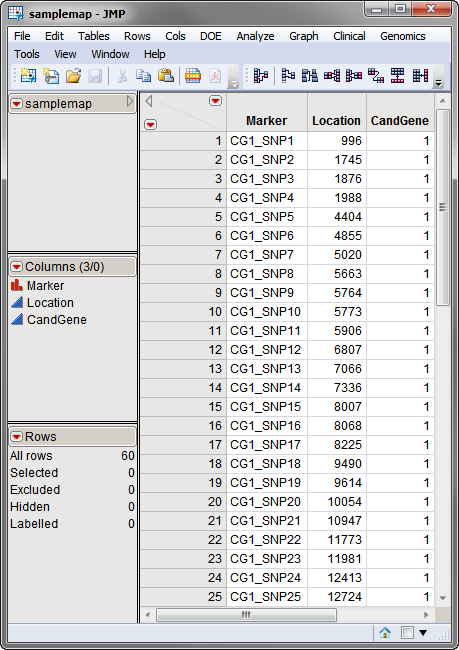
used in this example, the
samplemap.sas7bdat
data set, was computer generated and identifies markers, location and gene identities. A portion of this data set is illustrated below. This data set is a tall data set; each row corresponds to a different marker.
Note
: The top-to-bottom order of the rows in the annotation data set matches the left-to-right order of the columns in the input data set. This correspondence is required for markers to be matched appropriately.
For detailed information about the files and data sets used or created by JMP Life Sciences software, see
Files and Data Sets
.
The output generated by this process is summarized in a Tabbed report. Refer to the
Missing Genotype by Trait Summary
output documentation for detailed descriptions and guides to interpreting your results.