Genomic data sets often contain values or observations that deviate substantially from expected values. These deviations can often be ascribed to a variety of technical or other factors, that have little to do with biological cause and effects. In these situations, it is often better to identify and exclude outliers from an analysis than it is to include them, particularly where the cause of the anomalies can be explained by technical problems.
Often, valuable information about the quality of an array can be obtained from a visual examination of an image of the hybridization pattern. The Surface Summary process works with three dimensional data, with variables denoted X, Y, and Z. You can specify any number of Z variables to be plotted over common X and Y variables. The process summarizes the Z variables over gridded blocks of the X and Y variables and then creates a surface plot over the grid. You can optionally smooth the surface or subtract it from the original Z data. For example, you can use this process to perform spatial background correction for microarray data.
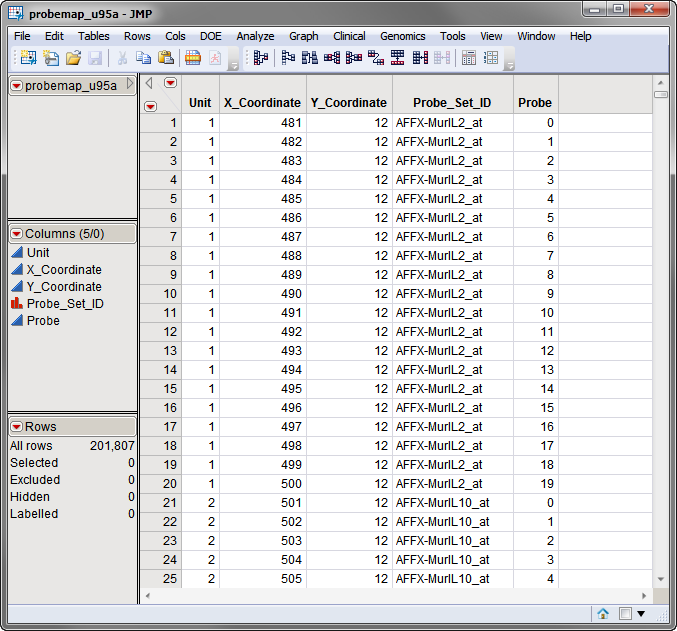
The first required data set, the coordinate data set, lists the x- and y-coordinates for each spot on the arrays. The probemap_u95a.sas7bdat data set, used in the following example, is shown below.
The second required data set, the input Z data set, contains all of the numeric Z value data (intensities) to be analyzed. The sample data set used in the following example, the chips45and55.sas7bdat data set, shown below, is from the Affymetrix Latin Square experiment that is described in Sample Case Studies. This data was originally generated by Affymetrix Corporation to develop and validate their U95A GeneChip and Microarray Suite (MAS) 5.0 algorithm over a range of known concentrations. Each group in the experiment contains a pool of non-specific RNA as well as a set of 14 distinct human transcripts spiked in at known concentrations. Intensity data are listed in the last two columns of the data set.
The probemap_u95a.sas7bdat coordinate data set and the chips45and55.sas7bdat input Z data set, are included in the Sample Data folder.
For detailed information about the files and data sets used or created by JMP Life Sciences software, see Files and Data Sets.
The output generated by this process is summarized in a Tabbed report. Refer to the Surface Summary output documentation for detailed descriptions and guides to interpreting your results.