The K Matrix Compression process clusters a symmetric relationship or estimated kinship matrix in order to reduce the size of the random effect that accounts for relatedness in the Q-K Mixed Model analytical process. The compressed matrix can be used to define the covariance structure in a down-stream mixed models for testing SNP-trait association. Compression can be done interactively through the JMP Clustering Platform where you can decide cluster membership via a cutoff level in hierarchical clustering based on visual inspection. Alternatively, clustering via PROC CLUSTER can be automated to produce a compressed K matrix based on a specified number of clusters. The optimized compression method scans through varying levels of compression to find the compression level that optimizes the fit of the mixed model to a specified trait (with the SNP effect excluded). This algorithm is described in Zhang et al. (Nature Genetics, 2010).
Warning: In contrast to the Q-K Mixed Model process, which requires the square root of the K matrix to be used in the model, the K Matrix Compression process requires the K matrix before taking the square root. This process computes the square root of the compressed K matrix (via Singular Value Decomposition) so that the columns are appropriately formatted for input as random effect variables in the Q-K Mixed Model process.
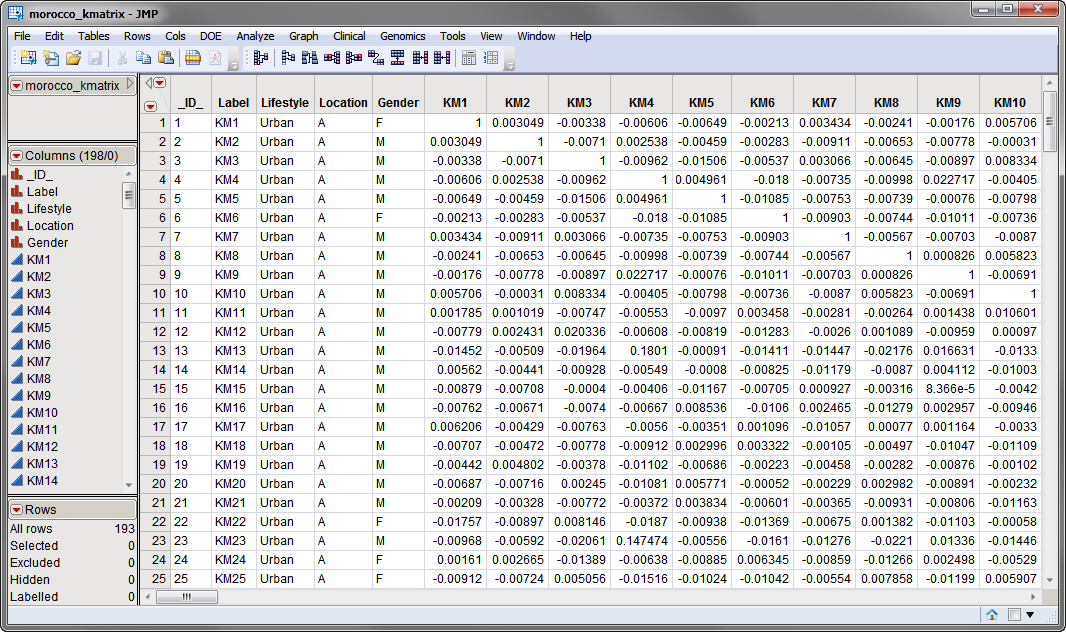
Two data sets are needed for this process. The first required data set is the input K matrix data set. The morocco_kmatrix.sas7bdat data set, shown below, represents a 193 x 193 matrix of Identity By Descent (IBD) values calculated between individuals from genome-wide SNP using data from a study of gene expression variation and SNP associations in southern Morocco (Idaghdour, Czika, et al., 2010)
The second data set is the SNP input data set. The SNP input data set used in the following example, the morocco_snps1exp.sas7bdat data set, lists gene expression variation data and SNP associations in southern Morocco (Idaghdour, Czika, et al., 2010).
Note: To run optimized compression, a trait variable must be specified from the SNP Input Data Set tab. If the SNP, phenotype data and K matrix are all in the Input K matrix data set, a separate SNP input data set is not required. However, the K matrix input data set should be specified again as the SNP Input Data.
The morocco_snps1exp.sas7bdat data set lists genotype data at 4744 SNPs in 193 individuals. Marker data is presented in the one-column format. This data set is partially shown below. Note that this is a wide data set; markers are listed in columns, whereas individuals are listed in rows.
Both data sets are included in the Sample Data folder.
For detailed information about the files and data sets used or created by JMP Life Sciences software, see Files and Data Sets.
The output generated by this process is summarized in a Tabbed report. Refer to the K Matrix Compression output documentation for detailed descriptions and guides to interpreting your results.