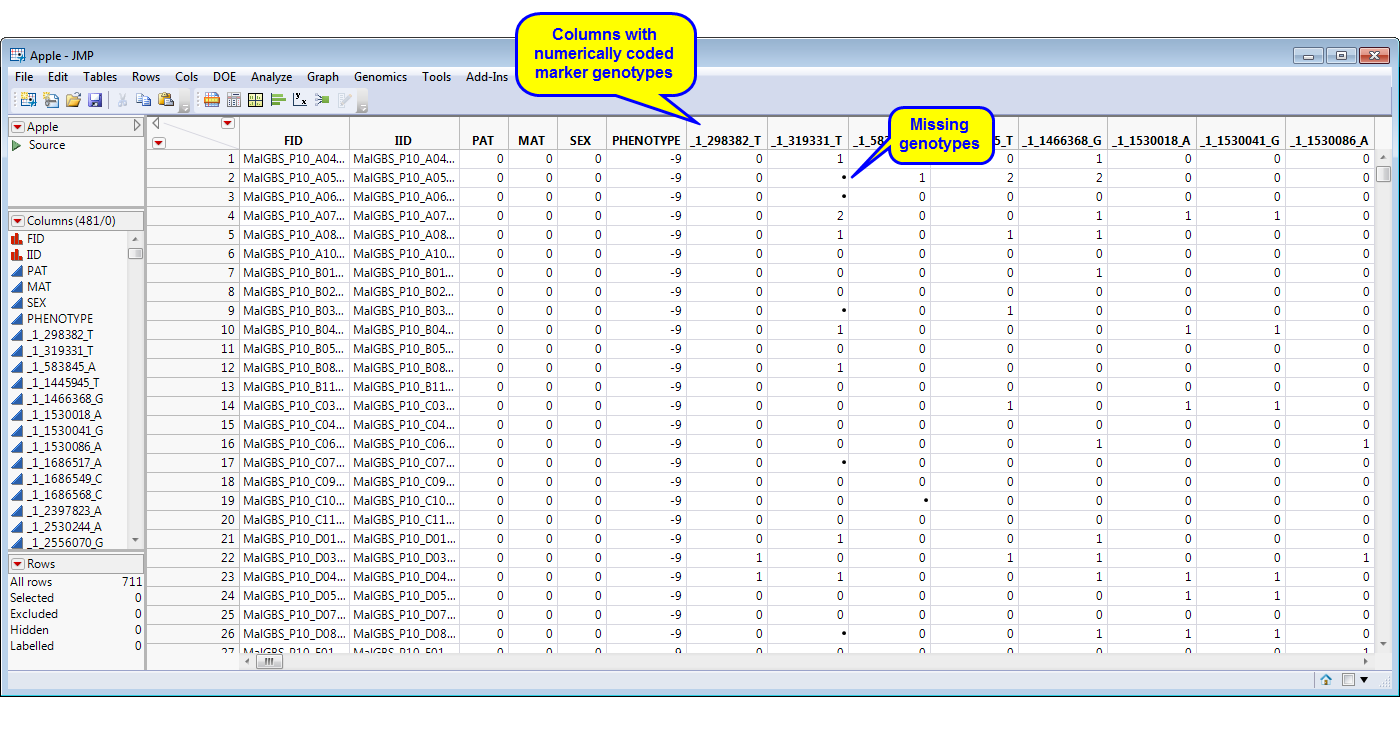
The Impute Missing Genotypes process imputes numeric missing marker genotypes (0, 1, or 2) for diploid organisms using the k-nearest neighbor imputation (kNNi) or the linkage disequilibrium k-nearest neighbor imputation (LD-kNNi) methods1. LD between markers is computed (using the SAS PROC ALLELE), distances between samples are computed (using the SAS PROC DISTANCE), and k-nearest neighbor samples is computed (using the SAS PROC MODECLUS)2.
One SAS data set is required: An input data set with one column per each numeric coded marker (0 for the homozygous major allele, 1 for the heterozygous, and 2 for the homozygous minor allele).
The second data set is the Annotation Data Set. This data set contains information, such as gene identity or chromosomal location, for each of the markers.
Output from this process is accessed from a Results window. Refer to the Impute Missing Genotypes output documentation for detailed descriptions and guides to interpreting your results.
Refer to the SAS PROC ALLELE, SAS PROC DISTANCE, and SAS PROC MODECLUS documentation for more information.