Stacked data sets typically have all of the variables of interest stacked into a single set of columns.
The Unstack process transposes a stacked data set into a tall SAS data set, splitting the variables of interest among multiple columns, and generates an accompanying Experimental Design Data Set (EDDS).
All you need to run this process is a stacked Input Data Set. A typical stacked data set is shown below.
Typically, stacked data sets contain a smaller number of columns when compared with the number of rows (circled above). This data set has 100300 data rows, but only 7 columns. All of the actual intensity data is stacked into the one log2i column! You should also note the repetition of the ChipID, Experiment, and Series columns for all probes on the array.
For detailed information about the files and data sets used or created by JMP Life Sciences software, see Files and Data Sets.
The output of the Unstack process includes the Results window, which lists the output data set and EDDS.
|
|
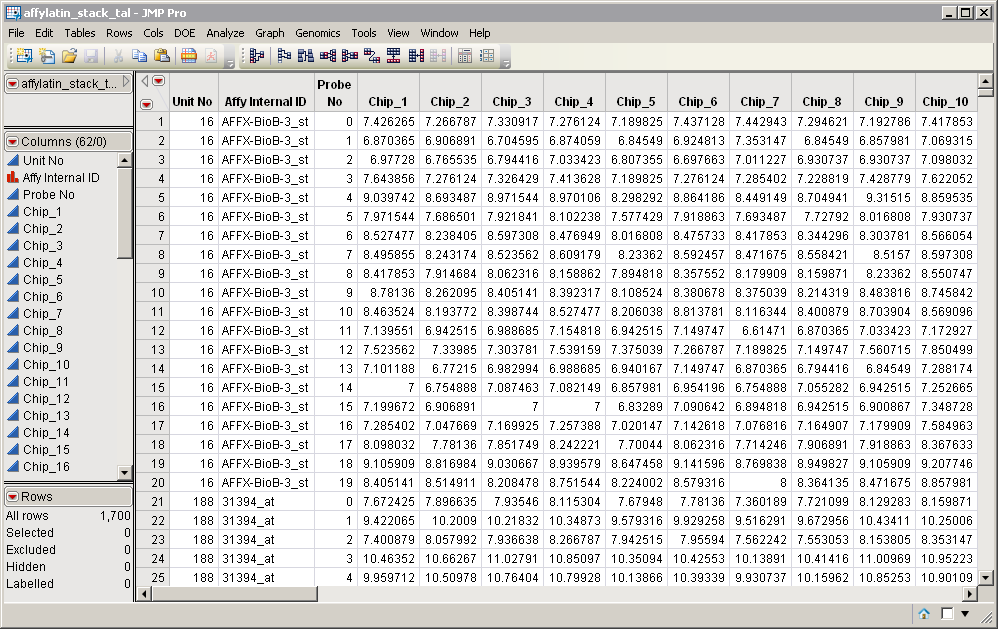
Click to view the affylatin_stack_tal.sas7bdat output data set.
|
The number of data columns in the output data set has increased to 59 from one in the input data set. This represents a 59-fold increase. The number of rows decreased from 100300 to 1700; a corresponding 59-fold decrease. All of the responses have been grouped into columns, with each column representing an individual array. Column names all contain the Chip_ prefix. The Series column has been eliminated.
|
|
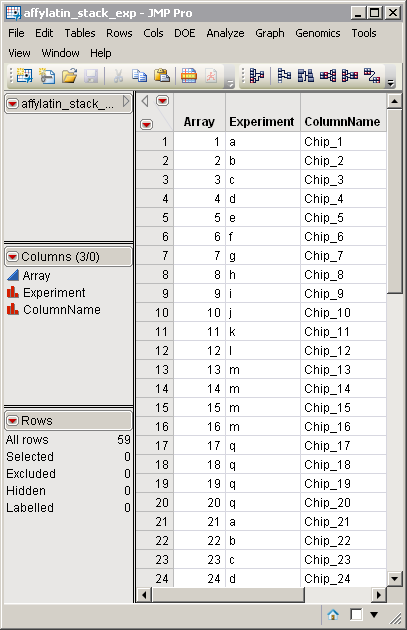
Click to view the affylatin_stack_exp.sas7bdat output EDDS.
|
|
•
|