Design That Estimates All Two-Factor Interactions
The Alias Matrix in Figure 5.5 shows partial aliasing of effects. In other cases, main effects might be fully aliased, or confounded, with two-factor interactions. In both of these cases, strong two-factor interactions can confuse the results of main effects only experiments. To avoid this risk, create a design that resolves all two-factor interactions.
In this example, you create a resolution V screening design. Two-factor interactions are orthogonal, but they are confounded with three-factor interactions.
1. Select DOE > Custom Design.
2. Type 5 next to Add N Factors.
3. Click Add Factor > Continuous.
4. Click Continue.
5. In the Model outline, select Interactions > 2nd.
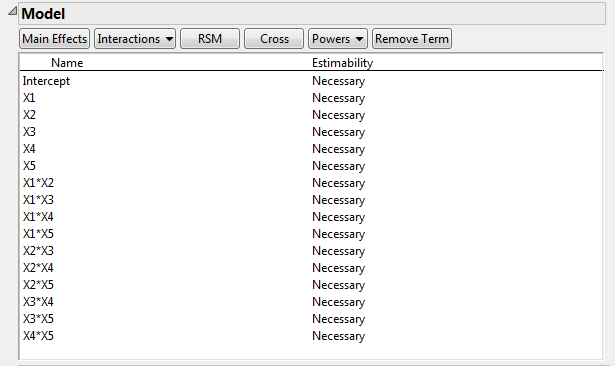
Figure 5.6 Model Outline Showing Interactions
6. Click Minimum to accept 16 for the number of runs.
Note: Setting the Random Seed in step 7 and Number of Starts in step 8 reproduces the exact results shown in this example. In constructing a design on your own, these steps are not necessary.
7. (Optional) Click the Custom Design red triangle, select Set Random Seed, type 12345, and click OK.
8. (Optional) Click the Custom Design red triangle, select Number of Starts, type 1, and click OK.
9. Click Make Design.
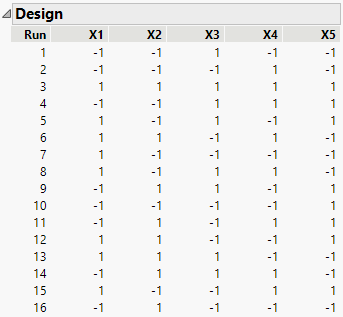
Figure 5.7 shows the runs of the design. All main effects and two-factor interactions are estimable because their Estimability was designated as Necessary (by default) in the Model outline.
Figure 5.7 Design to Estimate All Two-Factor Interactions
10. Open the Design Evaluation > Color Map on Correlations outline.
Figure 5.8 Color Map on Correlations
The Color Map indicates that the five main effects and the ten two-way interactions are all mutually orthogonal.