Publication date: 01/13/2026
Example of Distance Matrix
Learn how to compute various measures of distance or dissimilarity between observations in your life sciences genetic data.
1. Select Help > Sample Data Folder and open Iris.jmp.
2. Select Analyze > Life Sciences > Distance Matrix.
3. Select the first four columns (Sepal length, Sepal width, Petal length, and Petal width) and click Y, Columns.
4. Select Species and click X, Grouping Columns.
5. Click OK.
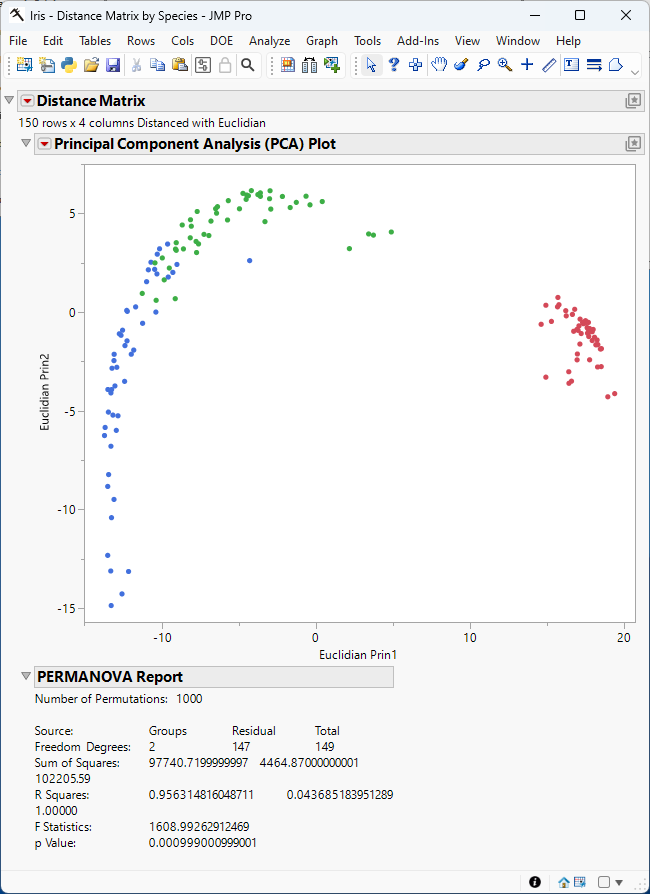
Figure 9.1 The Distance Matrix Report
Figure 9.1 shows the Principal Component Analysis (PCA) plot. Points that correspond to the observations in the input table are colored by species. The plot shows that the observations cluster by species and that the variance in sepal and petal length and width in irises is greatest between species.
Want more information? Have questions? Get answers in the JMP User Community (community.jmp.com).