Launch the Distance Matrix Platform
Launch the Distance Matrix platform by selecting Analyze > Life Sciences > Distance Matrix.
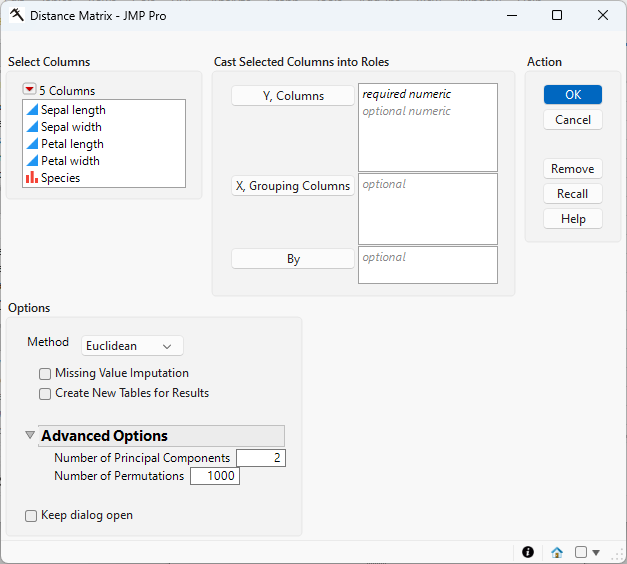
Figure 9.2 The Distance Matrix Launch Window
For information about the options in the Select Columns red triangle, see “Column Filter Menu” in Using JMP.
Y, Columns
Select the feature columns that you want, and click Y, Columns to specify the features that you want to consider in computing distances between observations or rows.
X, Grouping Columns
Specifies the column that defines groups for PERMANOVA analysis. Use this option to compute PERMANOVA results based on the selected grouping column, using the previously computed distance matrix.
By
Generates a separate report for each level of the By variable. If more than one By variable is assigned, a separate report is generated for each possible combination of their levels.
Method
Select the algorithm to use for performing the analysis.
• Select Euclidean to determine the straight-line distance between two points in Euclidean space. This method requires continuous data and is recommended for geometrically structured normalized data. This method is not ideal for use on sparse data or when variables are on different scales.
• Select Manhattan to compute the sum of the absolute differences across dimensions. This method is good for both continuous and ordinal data, especially when that data contains a grid-like structure. It is recommended for high-dimensional data and provides more robust handling of outliers than Euclidean methods.
• Select Gower when you have a mix of different types of data (continuous, ordinal, categorical, and binary). This method is recommended for sociological, ecological, or survey data. It is not ideal for large-scale pure numeric data.
• Select Bray-Curtis when you are measuring proportional differences rather than absolute differences. This method ignores shared zeros. The data should be nonnegative and continuous, consisting of counts or abundances. It is recommended for ecological data, microbiome data, species abundance data, and so on. It is not ideal for situations where the total magnitude is particularly meaningful.
• Select Jaccard to measure dissimilarity for sets or binary vectors. This method requires binary data, where there are only two possible answers. It is recommended for genetic data, text mining, or set comparisons. It should not be used when counts or magnitude matter.
• Select Binary to compute distance that is based on binary matching. This often implies a Simple Matching Coefficient. The data must be binary. This method is recommended for symmetrical binary attributes, where either result is equally important. It is not recommended for asymmetrical data, where one result is more significant than the other.
• Select Hamming to analyze the number of different positions between two equal-length strings. The data can be categorical, binary, or composed of sequences. It is used for analyzing DNA or protein sequences, binary strings, and error detection. It is recommended for categorical sequence comparisons but not suitable for continuous numeric data.
For details about each of these methods, see the SAS PROC Distance documentation.
Missing Value Imputation
Check this option when there are missing values to be imputed.
Create New Tables for Results
Check this option to create new tables for the output data and results. When this option is not checked, output is added to the end of the input data table.
Number of Principal Components
Use this option to specify the number of principal components to calculate. All of the specified principal components are added to the output table. Note that only the first two PCs are plotted.
Number of Permutations
Use this option to specify the number of permutations for computing PERMANOVA results.
Keep dialog open
Check this box to keep this platform window open after the analysis is run.
Data Format
Most of the processes in JMP assume that the input table has a particular data structure. JMP distinguishes between tall and wide data sets. A tall data table has samples as columns and molecular entity (for example, marker, gene, clone, protein, or metabolite) as rows, whereas a wide data table is the transpose of the tall data table, with samples as rows and molecular entities as columns.
When specifying the input data set for a process, it is important to know the required form. Distance Matrix requires a wide data table. The Transpose platform under the Tables menu enables you to transform your data tables between tall and wide forms.