Iterative Clustering Control Panel
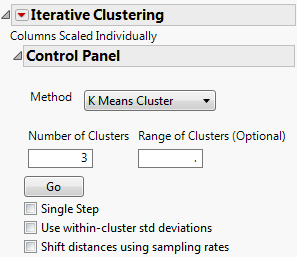
The Control Panel for the Cytometry.jmp data table is shown in Figure 13.7. You can iteratively fit different numbers of clusters or you can specify a range using the Range of Clusters option.
Figure 13.7 Iterative Clustering Control Panel
The Control Panel has the following options:
Method
The following clustering methods are available:
KMeans Clustering
Described in this chapter.
Self Organizing Map
Described in Self Organizing Map Control Panel.
Number of Clusters
Designates the number of clusters to form.
Range of Clusters (Optional)
Provides an upper bound for the number of clusters to form. If a number is entered here, the platform creates separate analyses for every integer between Number of Clusters and the value entered as Range of Clusters (Optional).
Go
Unless Single Step is selected, fits the clusters automatically.
Single Step
Enables you to step through the clustering process one iteration at a time. When you select Single Step and click Go, a K Means Cluster report appears with no cluster assignments but containing a Go and a Step button.
– Click the Step button to step through the iterations one at a time.
– Click the Go button to fit the clusters automatically.
Use within-cluster std deviations
Scales distances using the estimated standard deviation of each variable for observations within each cluster. If you do not select this option, distances are scaled by an overall estimate of the standard deviation of each variable.
Shift distances using sampling rates
Adjusts distances based on the sizes of clusters. If you have unequally sized clusters, an observation should have a higher probability of being assigned to larger clusters because there is a higher prior probability that the observation comes from a larger cluster.