Launch the Normalization Platform
Launch the Normalization platform by selecting Analyze > Life Sciences > Normalization.
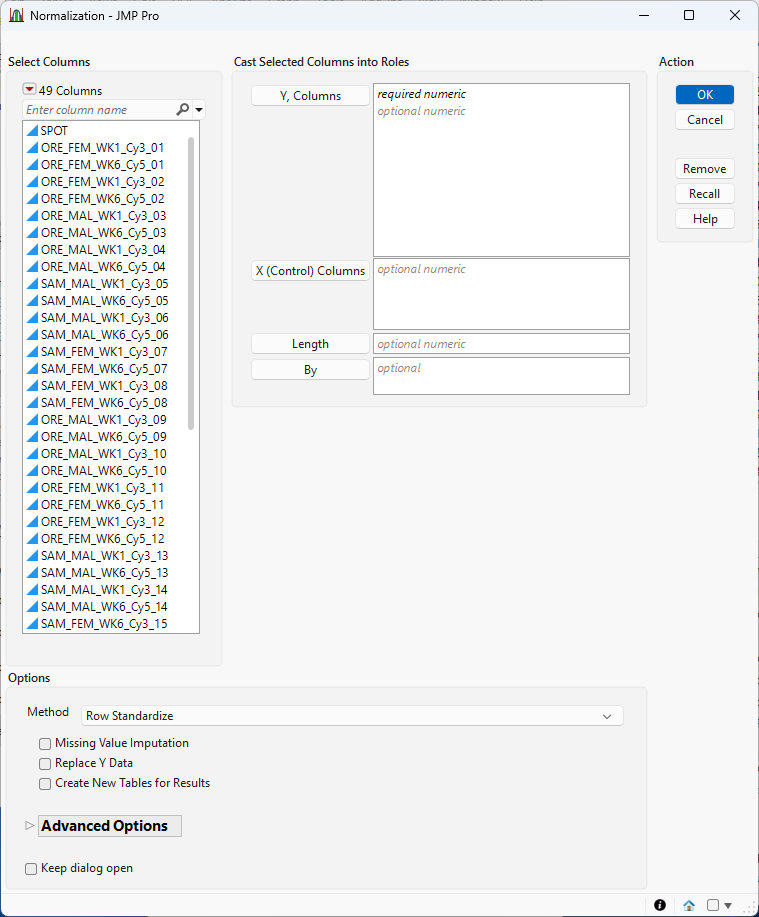
Figure 8.2 The Normalization Launch Window
For information about the options in the Select Columns red triangle, see “Column Filter Menu”in Using JMP.
Y, Columns
Select the sample columns that you want, and click Y, Columns to specify the samples that you want to analyze.
X, Control Columns
Use this option to specify known control or reference columns that contain one or more samples expected to represent a normal population. These help anchor normalization when using methods like Trimmed Mean of M-Values (TMM).
Length
Use this option to specify the column that contains the length of each transcript.
By
Generates a separate report for each level of the By variable. If multiple By variables are assigned, a report is generated for every possible combination of their levels.
Method
Select the algorithm to use for performing the normalization.
• Select Row Standardize to scale each row to have a mean of 0 and a standard deviation of 1. This method requires continuous data and is not ideal for compositional data or raw counts. The method is useful for highlighting relative variation within samples, particularly when combined with clustering and principal components analysis.
• Select Relative Abundance Percentage (RAP) to convert raw counts into percentage abundances across each sample. This method is used for counts data. It is used to emphasize relative proportions and should not be used for statistical models that are sensitive to compositional bias. It is particularly useful for visualizations that emphasize relative amounts, such as pie charts or bar graphs, and for exploratory analysis.
• Select Relative Abundance Ratio (RAR) to express each feature's abundance as a ratio relative to a reference (for example, a control or geometric mean). This method is used for counts data. It is used to highlight fold changes relative to a baseline. It is not ideal for either skewed or sparse data.
• Select Centered Log Ratio (CLR) to log-transform compositional data using the geometric mean as the center. This method is used to remove the closure effect in compositional data. It is particularly suited for microbiome and metagenomics data sets. This method is not recommended for data that contain ”zero” counts.
• Select Count/Read Per Million (CPM/RPM) to scale counts by total reads in each sample, and then multiply by one million. This method is used to adjust sequencing data to account for differences in sequencing depth when investigating expression comparisons across samples. This method is not recommended for data that contain ”zero” counts.
• Select Read/Fragment Per Kilobase of Transcript Per Million Mapped Reads/Fragments (RPKM/FPKM) to normalize sequencing data for both gene length and sequencing depth. This method is used for count data. It is particularly useful with transcriptomic expression profiling and between-sample comparisons. Because transcript length is considered, this method is recommended for between-sample comparisons.
• Select Transcript Per Million (TPM) to normalize sequencing data for both gene length and sequencing depth. This method is similar to RPKM, but is more suitable for between-sample comparisons. The method is used for count data, and reduces compositional bias in RNA-seq. It is recommended for differential expression (for example, edgeR) data, but should not be used for very sparse or low-depth data sets.
• Select Trimmed Mean of M-Values (TMM) to perform a normal or log-ratio normalization by using a reference sample that is trimmed to reduce bias. This method uses one or more reference or control samples when provided, or automatically selects a reference sample if none are specified. The method reduces compositional bias in RNA-seq and is recommended for differential expression analysis (for example, edgeR) data. It is not suitable for very sparse or heterogeneous data sets.
• Select Kernel Density Mean of M-Component (KDMM) to perform a normal or log-ratio density-aware normalization by using smoothed M-component distributions. This method takes weighted means of log-ratios across kernel-estimated density regions and adjusts for multimodal and nonlinear compositional structure. It can be applied to count or compositional data. It is recommended for advanced microbiome and metagenomics normalization, but should not be used for small sample sizes or low-resolution data.
Missing Value Imputation
Check this option when there are missing values to be imputed.
Replace Y-Data
Check this option to replace the original Y-columns with the corresponding normalized Y-columns.
Create New Tables for Results
Check this option to create new tables for the output data and results. When this option is not checked, output is added to the end of the input data table.
M Threshold
Use this advanced option to set a cutoff for log fold-change (M-values) between samples. This trims out extreme values that might distort normalization. Formula: M = log2(sample / reference). Using this option helps improve stability, especially in data sets with strong differential expression.
This option is used for TMM and KDMM only.
A Threshold
Use this advanced option to apply a lower limit to average log-intensity (A-values) so that low-abundance or noisy features do not skew the normalization. Formula: A = 0.5 × log2(sample × reference)
This option is used for TMM and KDMM only.
Prior Count
Use this advanced option to add a small constant to all counts before log transformation (for example, log2(count + 0.5)). This prevents issues with zeros in sparse data sets.
This option is used for TMM and KDMM only.
Interpolated Quantiles
Use this advanced option to enable the use of smoothed quantiles during normalization, which can be especially helpful in KDMM for complex or uneven data distributions.
This option is used for TMM and KDMM only.
Log Transformed
Use this advanced option to convert the raw counts to log-transformed values.
This option is used for TMM and KDMM only.
Keep dialog open
Check this box to keep this platform window open after the analysis is run.
Data Format
Most of the processes in JMP assume that the input table has a particular data structure. JMP distinguishes between tall and wide data tables. A tall data table has samples as columns and molecular features (for example, genes, transcripts, proteins, or metabolites) as rows, whereas a wide data table is the transpose of the tall data table, with samples as rows and molecular features as columns.
When specifying the input data set for the Normalization platform, it is important to know the required form. The Normalization platform expects a tall data table, where each row represents a molecular feature and each column represents a sample or condition. If your data are in the wide form, you can use the Transpose platform under the Tables menu to convert between tall and wide formats before normalization.
Normalization methods require numeric data values that represent measured or counted abundances (for example, read counts, intensities, or relative abundances). Nonnumeric or categorical data columns should be excluded from the normalization process.